runAS performs the following:
- Downloads one or more UniProt Taxonomic .dat files and creates FASTA files from them.
- Downloads the full SwissProt or TrEMBL, creates subset files by removing entries from the taxonomic
downloads, and creates a FASTA file of the remaining entries.
- Download the GO schema and data, and creates a GO mySQL database
with the mapping of UniProt IDs to the GO, KEGG, EC, Pfam and InterPro.
These are used for annotating a TCW database with
runSingleTCW.
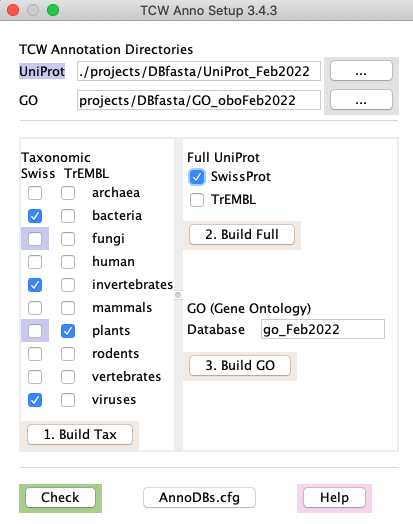
The highlighted entries in the figure indicate what has been created, i.e. the plant and fungi SwissProt.
When Build Tax is executed again, the 4 checked entries will be processed.
Other databases, such as Genbank nr, can also be used in runSingleTCW for annotation,
but the user must download and format them as FASTA files.
|
|


