|
FPC (FingerPrinted Contigs) | |||||||||||||||||||||||||||||||||||||||||
|
||||||||||||||||||||||||||||||||||||||||||
|
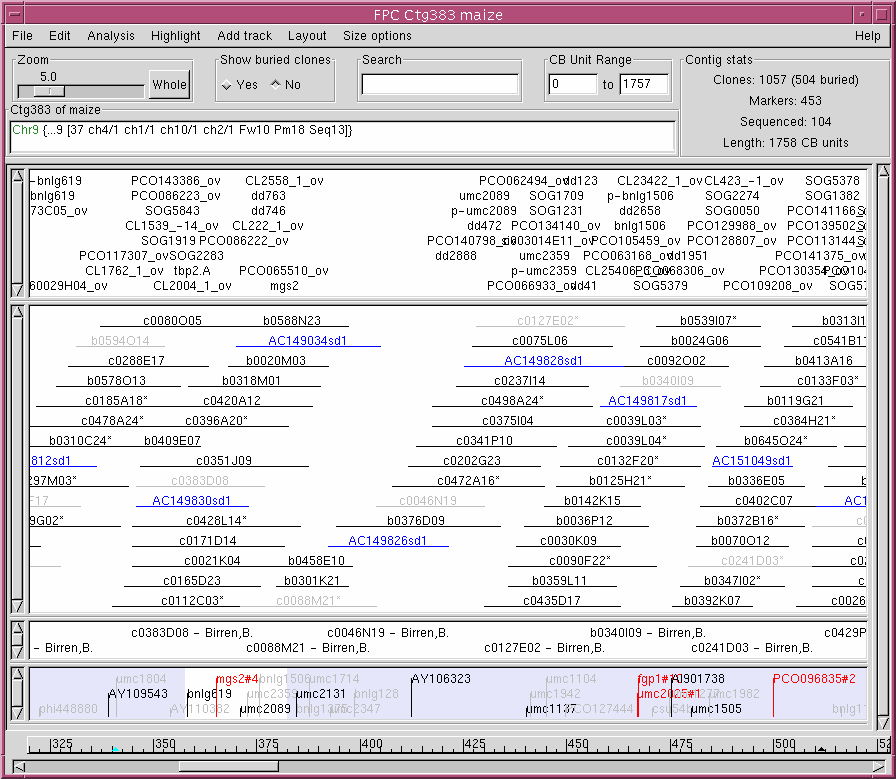
This work was funded in part by NSF grant #0213764 and by the Initiative for Future Agriculture and Food Systems Grant no. 2001-52100-11292 from the USDA Cooperative State Research, Education, and Extension Service. FPC takes as input a set of clones and their restriction fragments (called Bands) and assembles the clones into contigs; for more information, see summary. FPC contributions are by the GSC, BCGSC, Sanger Centre, Clemson University, and University of Arizona. Latest release: FPC V9.4 (21 Mar 2011) support for sequence based fingerprints.
FPC downloads, tutorials and documentation
FPC binaries, tutorials, documentation, and source code have been collected into a
single download package. The FPC Package contains the following (linked documents are viewable online):
*Published in Ian Dunham (ed) Genomic Mapping and Sequencing, Horizon Press, Genome Technology series. ReferencesNelson, W. and C. Soderlund (2009). Integrating Sequence with FPC Fingerprinted Maps. Nucleic Acids Research 1-11 doi:10.1093/nar/gkp034 Download Nelson, W., J. Dvorak, M. Luo, J. Messing, R. Wing, and C. Soderlund (2006). Efficacy of clone fingerprinting methodologies. Genomics 89:160-165. Download Nelson, W, A Bharti, E Butler, F Wei, G Fuks, H Kim, R Wing, J Messing, and C Soderlund (2005). Whole-Genome Validation of High-Information-Content Fingerprinting. Plant Physiology 139:27-38. Download Engler, F., J. Hatfield, W. Nelson, and C. Soderlund (2003). Locating sequence on FPC maps and selecting a minimal tiling path. Genome Research 13:2152:2163. Download Supplemental Soderlund, C., S. Humphrey, A. Dunhum, and L. French (2000). Contigs built with fingerprints, markers and FPC V4.7. Genome Research 10:1772-1787. Download. Soderlund, C., I. Longden and R. Mott (1997) FPC: a system for building contigs from restriction fingerprinted clones. CABIOS, 13: 523-535. Download. Book chapters Nelson, W. and C. Soderlund (2005). Software for restriction fragment physical maps. K. Meksem, G. Kahl (ed) The Handbook of Genome Mapping: Genetic and Physical Mapping, Wiley-VCH. pp. 285-306. Download Engler, F. and C. Soderlund (2002). Software for Physical Maps. In Ian Dunham (ed) Genomic Mapping and Sequencing, Horizon Press, Genome Technology series. Norfolk, UK, pp. 201-236. Soderlund, C., S. Gregory and I. Dunham (1998). Sequence ready clones. In Bishop, M.J. (ed) Guide to Human Genome Computing, Academic Press. Gregory, S., C. Soderlund and A. Coulson (1996) Contig assembly by fingerprinting. In P. H. Dear (ed) Genome Mapping: a Practical Approach. Other van Oeveren J, de Ruiter M, Jesse T, van der Poel H, Tang J, Yalcin F, Janssen A, Volpin H, Stormo KE, Bogden R, van Eijk MJ, Prins M (2011). Sequence-based physical mapping of complex genomes by whole genome profiling. Genome Res., epub Feb. 1, 2011. Download Ness, S.R., Terpstra, W., Krzywinski, M., Marra M., Jones, A., and Ness, S.J.M. (2002). Assembly of fingerprint contigs: parallelized FPC. BioInformatics 18:484-485. The International Human Genome Mapping Consortium (2001) A physical map of the human genome, Nature 409:934-941 Download. | ||||||||||||||||||||||||||||||||||||||||||
| Email Comments To: fpc@agcol.arizona.edu |